MBD0042
Escherichia coli FISH probe - Cy3
Probe for fluorescence in situ hybridization (FISH)
Přihlásitk zobrazení cen stanovených pro organizaci a smluvních cen
About This Item
UNSPSC Code:
12352200
NACRES:
NA.55
Doporučené produkty
Quality Level
technique(s)
FISH: suitable
fluorescence
λex 550 nm; λem 570 nm
shipped in
dry ice
storage temp.
−20°C
General description
Fluorescent In Situ Hybridization technique (FISH) is based on the hybridization of fluorescent labeled oligonucleotide probe to a specific complementary DNA or RNA sequence in whole and intact cells.1 Microbial FISH allows the visualization, identification and isolation of bacteria due to recognition of ribosomal RNA also in unculturable samples.2
FISH technique can serve as a powerful tool in the microbiome research field by allowing the observation of native microbial populations in diverse microbiome environments, such as samples from human origin (blood3 and tissue4), microbial ecology (solid biofilms5 and aquatic systems 6) and plants7. It is strongly recommended to include positive and negative controls in FISH assays to ensure specific binding of the probe of interest and appropriate protocol conditions. We offer positive (MBD0032/33) and negative control (MBD0034/35) probes, that accompany the specific probe of interest.
Escherichia coli is a gram negative, facultative aerobic, rod-shaped coliform bacterium. E. coli colonizes the infant gut within hours of birth and establishes itself as the most abundant facultative anaerobe of the human intestinal microflora for the remainder of life, equipped with the abilities to grow in the ever-changing environment in the gut and cope with the mammalian host interaction.8,9 Nevertheless, E. coli can survive in many different ecological habitats, including abiotic environments, and is considered a highly versatile species. Known habitats of E. coli include soil, water, sediment, and food. Some strains of E. coli have evolved and adapted to a pathogenic lifestyle and can cause different disease pathologies.10
Escherichia coli probe specifically recognizes Escherichia coli cells.
Yet there are some reports that describe recognition of other bacteria with this probe, such as, Shigella boydii, Citrobacter davisae, Citrobacter lapagei, Citrobacter neteri 11 and Klebsiella pneumoniae 12.
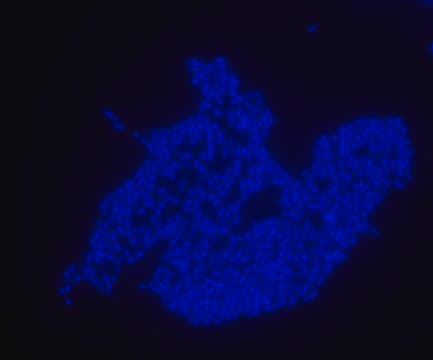
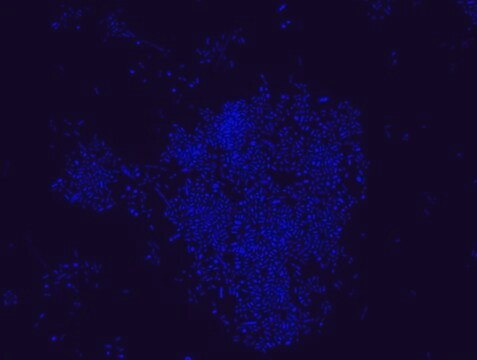
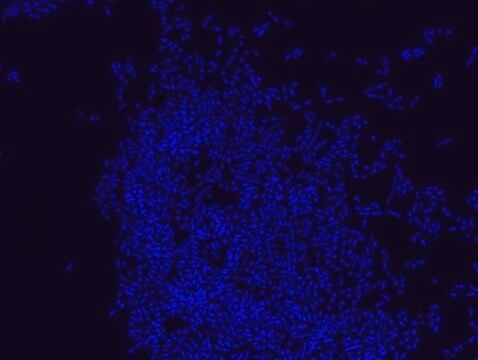
FISH technique was successfully used to identify E.coli with the probe in various samples such as pure culture (as described in the figure legends and11,13), large and small intestines samples14-16, fecal samples17-21, colonic biopsies18, urine samples, bladder and kidney sections embedded in paraffin22 and in E.coli biofilm23.
FISH technique can serve as a powerful tool in the microbiome research field by allowing the observation of native microbial populations in diverse microbiome environments, such as samples from human origin (blood3 and tissue4), microbial ecology (solid biofilms5 and aquatic systems 6) and plants7. It is strongly recommended to include positive and negative controls in FISH assays to ensure specific binding of the probe of interest and appropriate protocol conditions. We offer positive (MBD0032/33) and negative control (MBD0034/35) probes, that accompany the specific probe of interest.
Escherichia coli is a gram negative, facultative aerobic, rod-shaped coliform bacterium. E. coli colonizes the infant gut within hours of birth and establishes itself as the most abundant facultative anaerobe of the human intestinal microflora for the remainder of life, equipped with the abilities to grow in the ever-changing environment in the gut and cope with the mammalian host interaction.8,9 Nevertheless, E. coli can survive in many different ecological habitats, including abiotic environments, and is considered a highly versatile species. Known habitats of E. coli include soil, water, sediment, and food. Some strains of E. coli have evolved and adapted to a pathogenic lifestyle and can cause different disease pathologies.10
Escherichia coli probe specifically recognizes Escherichia coli cells.
Yet there are some reports that describe recognition of other bacteria with this probe, such as, Shigella boydii, Citrobacter davisae, Citrobacter lapagei, Citrobacter neteri 11 and Klebsiella pneumoniae 12.
FISH technique was successfully used to identify E.coli with the probe in various samples such as pure culture (as described in the figure legends and11,13), large and small intestines samples14-16, fecal samples17-21, colonic biopsies18, urine samples, bladder and kidney sections embedded in paraffin22 and in E.coli biofilm23.
Application
Probe for fluorescence in situ hybridization (FISH),recognizes Escherichia coli cells
Features and Benefits
- Visualize, identify and isolate Escherichia coli cells.
- Observe native E. coli cell populations in diverse microbiome environments.
- Specific, sensitive and robust identification of E. coli in bacterial mixed population.
- Specific, sensitive and robust identification even when E. coli is in low abundance in the sample.
- FISH can complete PCR based detection methods by avoiding contaminant bacteria detection.
- Provides information on E.coli morphology and allows to study biofilm architecture.
- Identify E.coli in clinical samples such as, urine samples, bladder and kidney sections (formalin-fixed paraffin-embedded (FFPE) samples), fecal samples and colon tissue.
- The ability to detect E.coli in its natural habitat is an essential tool for studying host-microbiome interaction.
Storage Class
12 - Non Combustible Liquids
wgk_germany
nwg
flash_point_f
Not applicable
flash_point_c
Not applicable
Osvědčení o analýze (COA)
Vyhledejte osvědčení Osvědčení o analýze (COA) zadáním čísla šarže/dávky těchto produktů. Čísla šarže a dávky lze nalézt na štítku produktu za slovy „Lot“ nebo „Batch“.
Již tento produkt vlastníte?
Dokumenty související s produkty, které jste v minulosti zakoupili, byly za účelem usnadnění shromážděny ve vaší Knihovně dokumentů.
S Favre-Bonté et al.
Infection and immunity, 67(11), 6152-6156 (1999-10-26)
The role of the Klebsiella pneumoniae capsular polysaccharide (K antigen) during colonization of the mouse large intestine was assessed with wild-type K. pneumoniae LM21 and its isogenic capsule-defective mutant. When bacterial strains were fed alone to mice, the capsulated bacteria
L K Poulsen et al.
Infection and immunity, 62(11), 5191-5194 (1994-11-01)
Fluorescent oligonucleotide probes targeting rRNA were used to develop an in situ hybridization technique by which the spatial distribution of Escherichia coli in the large intestines of streptomycin-treated mice was determined. Single E. coli cells were identified in thin frozen
D Grimm et al.
Applied and environmental microbiology, 64(7), 2686-2690 (1998-07-02)
Based on comparative sequence analysis, we have designed an oligonucleotide probe complementary to a region of 16S rRNA of Legionella pneumophila which allows the differentiation of L. pneumophila from other Legionella species without cultivation. The specificity of the new probe
Arthur C Ouwehand et al.
Microbiology and immunology, 48(7), 497-500 (2004-07-24)
The fecal and mucosal microbiota of infants with rectal bleeding and the fecal microbiota of healthy age-matched controls were investigated by fluorescent in situ hybridization. Bifidobacteria were the main genus in both the feces and mucosa. The other genera tested
Stuart C Smith et al.
European journal of nutrition, 45(6), 335-341 (2006-06-10)
Changes in the composition of gastrointestinal microbiota by dietary interventions using pro- and prebiotics provide opportunity for improving health and preventing disease. However, the capacity of lupin kernel fiber (LKFibre), a novel legume-derived food ingredient, to act as a prebiotic
Náš tým vědeckých pracovníků má zkušenosti ve všech oblastech výzkumu, včetně přírodních věd, materiálových věd, chemické syntézy, chromatografie, analytiky a mnoha dalších..
Obraťte se na technický servis.