07-354
Anti-acetyl-Histone H3 (Lys18) Antibody
serum, Upstate®
Synonym(s):
H3K18Ac, Histone H3 (acetyl K18)
About This Item
Recommended Products
biological source
rabbit
Quality Level
antibody form
serum
antibody product type
primary antibodies
clone
polyclonal
species reactivity
yeast, Saccharomyces cerevisiae, human
species reactivity (predicted by homology)
vertebrates (most common)
manufacturer/tradename
Upstate®
technique(s)
ChIP: suitable (ChIP-seq)
dot blot: suitable
western blot: suitable
NCBI accession no.
UniProt accession no.
shipped in
dry ice
target post-translational modification
acetylation (Lys18)
Gene Information
human ... H3C1(8350)
General description
Acetylation of histone H3 occurs at several different lysine positions in the histone tail and is performed by a family of enzymes known as Histone Acetyl Transferases (HATs).
Specificity
Immunogen
application
Representative lot data.
Sonicated chromatin prepared from HeLa cells (1 X 10E6 cell equivalents per IP) were subjected to chromatin immunoprecipitation using 2 µL of either Normal Rabbit Serum , or 2 µL of Anti-Acetyl-Histone H3 (Lys18)and the Magna ChIP A Kit (Cat. # 17-610). Successful immunoprecipitation of Acetyl-Histone H3 (Lys18) associated DNA fragments was verified by qPCR using Control Primers specific for the human GAPDH promoter region as a positive locus, and β-globin primers as a negative locus. Data is presented as percent input of each IP sample relative to input chromatin for each amplicon and ChIP sample as indicated.
Please refer to the EZ-Magna ChIP A (Cat. # 17-408) or EZ-ChIP (Cat. # 17-371) protocol for experimental details.
ChIP-Sequencing:
Representative lot data. Chromatin immunoprecipitation was performed using the Magna ChIP HiSens kit (cat# 17-10460), 2 μg of anti-acetyl-Histone H3 (Lys18) antibody (cat# 07-354), 20 µL Protein A/G beads , and 1e6 crosslinked HeLa cell chromatin followed by DNA purification using magnetic beads. Libraries were prepared from Input and ChIP DNA samples using standard protocols with Illumina barcoded adapters, and analyzed on Illumina HiSeq instrument. An excess of ten million reads from FastQ files were mapped using Bowtie (http://bowtie-bio.sourceforge.net/manual.shtml) following TagDust (http://genome.gsc.riken.jp/osc/english/dataresource/) tag removal. Peaks were identified using MACS (http://luelab.dfci.harvard.edu/MACS/), with peaks and reads visualized as a custom track in UCSC Genome Browser (http://genome.ucsc.edu) from BigWig and BED files.
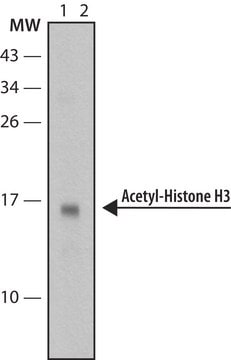
Western Blot Analysis:
Representative lot data.
Recombinant histone H3 (lane 1, Catalog # 14-494) and acid extracts from sodium butyrate treated (lane 2) and untreated (lane 3) HeLa cells (Catalog # 17-305) were probed with anti acetyl- Histone H3 (Lys18) (1:10,000 dilution).
Arrow indicates acetyl histone H3 (Lys18) (17 kDa).
Dot Blot:
Representative lot data.
40 ng and 4ng amounts of histone peptides with various modifications (see table 1) were transferred to PVDF membrane and probed with Anti-Acetyl-Histone H3 (Lys18) antibody (1:5000 dilution). Proteins were visualized using a goat anti-rabbit IgG conjugated to HRP and a chemiluminescence detection system. Image from a 60 second exposure is shown.
Quality
Target description
Linkage
Physical form
Other Notes
Legal Information
Not finding the right product?
Try our Product Selector Tool.
Storage Class
12 - Non Combustible Liquids
wgk_germany
WGK 1
flash_point_f
Not applicable
flash_point_c
Not applicable
Certificates of Analysis (COA)
Search for Certificates of Analysis (COA) by entering the products Lot/Batch Number. Lot and Batch Numbers can be found on a product’s label following the words ‘Lot’ or ‘Batch’.
Already Own This Product?
Find documentation for the products that you have recently purchased in the Document Library.
Our team of scientists has experience in all areas of research including Life Science, Material Science, Chemical Synthesis, Chromatography, Analytical and many others.
Contact Technical Service