推薦產品
mp
153-157 °C (lit.)
SMILES 字串
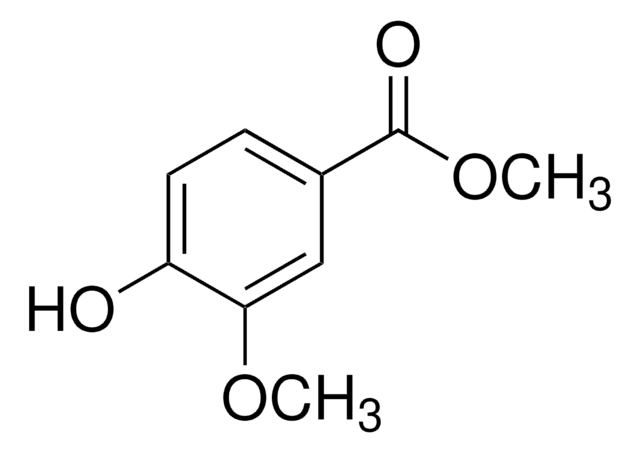
OC(=O)c1cccc(Cl)c1
InChI
1S/C7H5ClO2/c8-6-3-1-2-5(4-6)7(9)10/h1-4H,(H,9,10)
InChI 密鑰
LULAYUGMBFYYEX-UHFFFAOYSA-N
尋找類似的產品? 前往 產品比較指南
取代透過
產品號碼
描述
訂價
訊號詞
Warning
危險聲明
危險分類
Eye Irrit. 2 - Skin Irrit. 2
儲存類別代碼
11 - Combustible Solids
水污染物質分類(WGK)
WGK 3
閃點(°F)
Not applicable
閃點(°C)
Not applicable
個人防護裝備
dust mask type N95 (US), Eyeshields, Gloves
分析證明 (COA)
輸入產品批次/批號來搜索 分析證明 (COA)。在產品’s標籤上找到批次和批號,寫有 ‘Lot’或‘Batch’.。
Alfredo Gallego et al.
World journal of microbiology & biotechnology, 28(3), 1245-1252 (2012-07-19)
An indigenous strain of Pseudomonas putida capable of degrading 3-chlorobenzoic acid as the sole carbon source was isolated from the Riachuelo, a polluted river in Buenos Aires. Aerobic biodegradation assays were performed using a 2-l microfermentor. Biodegradation was evaluated by
Sudip K Samanta et al.
Molecular microbiology, 55(4), 1151-1159 (2005-02-03)
Rhodopseudomonas palustris strain RCB100 degrades 3-chlorobenzoate (3-CBA) anaerobically. We purified from this strain a coenzyme A ligase that is active with 3-CBA and determined its N-terminal amino acid sequence to be identical to that of a cyclohexanecarboxylate-CoA ligase encoded by
V P Jayachandran et al.
Journal of industrial microbiology & biotechnology, 36(2), 219-227 (2008-10-23)
The compatibility and efficiency of two ortho-cleavage pathway-following pseudomonads viz. the 3-chlorobenzoate (3-CBA)-degrader, Pseudomonas aeruginosa 3mT (3mT) and the phenol-degrader, P. stutzeri SPC-2 (SPC-2) in a mixed culture for the degradation of these substrates singly and simultaneously in mixtures was
Iris Plumeier et al.
Journal of bacteriology, 184(15), 4054-4064 (2002-07-11)
The tfdC(I)D(I)E(I)F(I,) and tfdD(II)C(II)E(II)F(II) gene modules of plasmid pJP4 of Ralstonia eutropha JMP134 encode complete sets of functional enzymes for the transformation of chlorocatechols into 3-oxoadipate, which are all expressed during growth on 2,4-dichlorophenoxyacetate (2,4-D). However, activity of tfd(I)-encoded enzymes
Caroline Laemmli et al.
Archives of microbiology, 181(2), 112-121 (2003-12-17)
Ralstonia eutropha JMP134 possesses two sets of similar genes for degradation of chloroaromatic compounds, tfdCDEFB (in short: tfdI cluster) and tfdDII CII EII FII BII (tfdII cluster). The significance of two sets of tfd genes for the organism has long
我們的科學家團隊在所有研究領域都有豐富的經驗,包括生命科學、材料科學、化學合成、色譜、分析等.
聯絡技術服務