MBD0032
Eubacteria FISH probe - ATTO488
Probe for fluorescence in situ hybridization (FISH), 20 µM in water
Synonym(s):
Fluorescent Probe
Sign Into View Organizational & Contract Pricing
All Photos(18)
About This Item
UNSPSC Code:
12352200
NACRES:
NA.55
Recommended Products
Quality Level
technique(s)
FISH: suitable
fluorescence
λex 504 nm; λem 521 nm (ATTO488)
shipped in
dry ice
storage temp.
−20°C
General description
Fluorescent In Situ Hybridization technique (FISH) is based on the hybridization of fluorescent labeled oligonucleotide probe to a specific complementary DNA or RNA sequence in whole and intact cells. Microbial FISH allows the visualization, identification and isolation of bacteria due to recognition of ribosomal RNA also in unculturable samples.
FISH technique can serve as a powerful tool in the microbiome research field by allowing the observation of native microbial populations in diverse microbiome environments, such as samples from human origin (blood and tissue), microbial ecology (solid biofilms and aquatic systems) and plants.
Prokaryotic single cell life forms are divided into two domains, called Bacteria and Archaea, originally categorized as Eubacteria and Archaebacteria. However both terms, Eubacteria and Bacteria are still being used in microbiology. Eubacteria probe recognizes most bacteria as it is complementary to a portion of 16S rRNA found in almost all bacteria.,
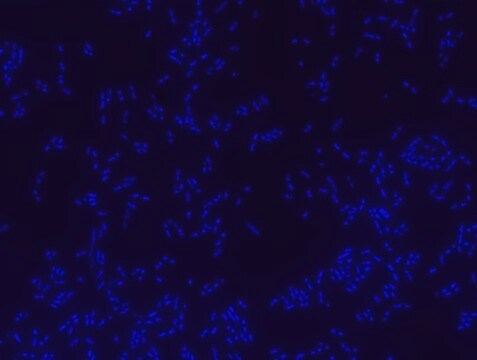
FISH technique was successfully used to identify different bacteria with the universal bacterial probe in various samples such as, pure culture (as described in the figure legends), blood cultures,, periapical tooth lesions12, saliva13, biofilms from voice prostheses14, subgingival biofilm15, aortic wall tissue16, buccal epithelial cells, pure culture and cell culture17, intestine tissue embedded in paraffin18, necrotizing fasciitis and pure culture19, colon sections embedded in paraffin20,21, cancer tissues22,23, environmental samples24 and gut of the medicinal leech25. The probe can also be used for combined technique of FISH and Flow cytometric analysis. 9,26,27
It is strongly recommended to include positive and negative controls in FISH assays to ensure specific binding of the probe of interest and appropriate protocol conditions. We offer positive (MBD0032/33) and negative (MBD0034/35) control probes, that accompany the specific probe of interest.
FISH technique can serve as a powerful tool in the microbiome research field by allowing the observation of native microbial populations in diverse microbiome environments, such as samples from human origin (blood and tissue), microbial ecology (solid biofilms and aquatic systems) and plants.
Prokaryotic single cell life forms are divided into two domains, called Bacteria and Archaea, originally categorized as Eubacteria and Archaebacteria. However both terms, Eubacteria and Bacteria are still being used in microbiology. Eubacteria probe recognizes most bacteria as it is complementary to a portion of 16S rRNA found in almost all bacteria.,
FISH technique was successfully used to identify different bacteria with the universal bacterial probe in various samples such as, pure culture (as described in the figure legends), blood cultures,, periapical tooth lesions12, saliva13, biofilms from voice prostheses14, subgingival biofilm15, aortic wall tissue16, buccal epithelial cells, pure culture and cell culture17, intestine tissue embedded in paraffin18, necrotizing fasciitis and pure culture19, colon sections embedded in paraffin20,21, cancer tissues22,23, environmental samples24 and gut of the medicinal leech25. The probe can also be used for combined technique of FISH and Flow cytometric analysis. 9,26,27
It is strongly recommended to include positive and negative controls in FISH assays to ensure specific binding of the probe of interest and appropriate protocol conditions. We offer positive (MBD0032/33) and negative (MBD0034/35) control probes, that accompany the specific probe of interest.
Application
Eubacteria FISH probe - ATTO488 is suitable to use as a probe for fluorescence in situ hybridization (FISH) to recognize Eubacteria cells .
Features and Benefits
- Visualize, identify and isolate bacteria cells.
- Observe native bacteria cell populations in diverse microbiome environments.
- Specific, sensitive and robust identification of bacteria cells in mixed microorganism population.
- Specific, sensitive and robust identification even when bacteria are in low abundance in the sample.
- FISH can complete PCR based detection methods by avoiding contaminant bacteria detection.
- Provides information on bacteria morphology and allows to study biofilm architecture.
- Identify various bacteria in environmental and clinical samples such as, formalin-fixed paraffin-embedded (FFPE) samples, blood cultures, saliva and more.
- The ability to detect bacteria in its natural habitat is an essential tool for studying host-microbiome interaction.
Storage Class Code
12 - Non Combustible Liquids
WGK
nwg
Flash Point(F)
Not applicable
Flash Point(C)
Not applicable
Certificates of Analysis (COA)
Search for Certificates of Analysis (COA) by entering the products Lot/Batch Number. Lot and Batch Numbers can be found on a product’s label following the words ‘Lot’ or ‘Batch’.
Already Own This Product?
Find documentation for the products that you have recently purchased in the Document Library.
Jiahui Yu et al.
International journal of cancer, 139(6), 1318-1326 (2016-05-01)
The prevalence of invasive Fusobacterium nucleatum (Fn) within the serrated neoplasia pathway of the proximal colon has seldom been investigated. We examined the invasive Fn and bacterial biofilms in 35 proximal hyperplastic polyps (HPs), 33 sessile serrated adenomas (SSAs), 48
K Trebesius et al.
Medical microbiology and immunology, 188(4), 169-175 (2000-08-05)
Fluorescence in situ hybridisation (FISH) targeted to ribosomal RNA is well established for studies in environmental microbiology. Initial applications of this technique in the field of medical microbiology showed that FISH is also a suitable means for the rapid, reliable
Michele A Maltz et al.
Frontiers in microbiology, 5, 151-151 (2014-05-27)
There are trillions of microbes found throughout the human body and they exceed the number of eukaryotic cells by 10-fold. Metagenomic studies have revealed that the majority of these microbes are found within the gut, playing an important role in
V A Kempf et al.
Journal of clinical microbiology, 38(2), 830-838 (2000-02-03)
Using fluorescent in situ hybridization (FISH) with rRNA-targeted fluorescently labelled oligonucleotide probes, pathogens were rapidly detected and identified in positive blood culture bottles without cultivation and biotyping. In this study, 115 blood cultures with a positive growth index as determined
Dorothee Maria Gescher et al.
International journal of antimicrobial agents, 32 Suppl 1, S51-S59 (2008-08-23)
Sepsis is a life-threatening disease with a high mortality rate. Rapid identification of blood culture isolates plays a crucial role in adequate antimicrobial therapy in sepsis patients. To accelerate microbiological diagnosis, a comprehensive panel of oligonucleotide probes for fluorescence in
Our team of scientists has experience in all areas of research including Life Science, Material Science, Chemical Synthesis, Chromatography, Analytical and many others.
Contact Technical Service