746223
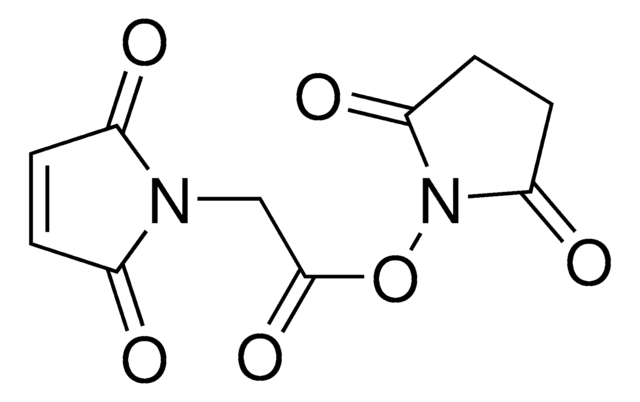
Maleimide-PEG2-succinimidyl ester
≥95%
Synonym(s):
3-[2-[2-[[3-(2,5-Dihydro-2,5-dioxo-1H-pyrrol-1-yl)-1-oxopropyl]amino]ethoxy]ethoxy]propanoic acid 2,5-dioxo-1-pyrrolidinyl ester, Maleimide-PEG2-NHS
About This Item
Recommended Products
Assay
≥95%
form
solid
reaction suitability
reaction type: Pegylations
reagent type: cross-linking reagent
mp
92-94 °C
functional group
NHS ester
maleimide
storage temp.
−20°C
SMILES string
O=C(N1CCC(NCCOCCOCCC(ON(C(CC2)=O)C2=O)=O)=O)C=CC1=O
InChI
1S/C18H23N3O9/c22-13(5-8-20-14(23)1-2-15(20)24)19-7-10-29-12-11-28-9-6-18(27)30-21-16(25)3-4-17(21)26/h1-2H,3-12H2,(H,19,22)
InChI key
TZPDZOJURBVWHS-UHFFFAOYSA-N
Related Categories
Application
Storage Class Code
11 - Combustible Solids
WGK
WGK 3
Flash Point(F)
Not applicable
Flash Point(C)
Not applicable
Certificates of Analysis (COA)
Search for Certificates of Analysis (COA) by entering the products Lot/Batch Number. Lot and Batch Numbers can be found on a product’s label following the words ‘Lot’ or ‘Batch’.
Already Own This Product?
Find documentation for the products that you have recently purchased in the Document Library.
Customers Also Viewed
Our team of scientists has experience in all areas of research including Life Science, Material Science, Chemical Synthesis, Chromatography, Analytical and many others.
Contact Technical Service




![O-[N-(3-Maleimidopropionyl)aminoethyl]-O′-[3-(N-succinimidyloxy)-3-oxopropyl]heptacosaethylene glycol ≥90% (oligomer purity)](/deepweb/assets/sigmaaldrich/product/structures/367/864/1c31a65f-501f-4464-8bb3-edff0d131a12/640/1c31a65f-501f-4464-8bb3-edff0d131a12.png)