MAC4L
MetaPolyzyme
lyophilized powder
Synonym(s):
Multilytic Enzyme Mix
About This Item
Recommended Products
form
lyophilized powder
Quality Level
technique(s)
DNA extraction: suitable
suitability
suitable for microbiology
application(s)
microbiology
shipped in
wet ice
storage temp.
−20°C
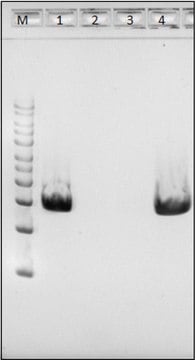
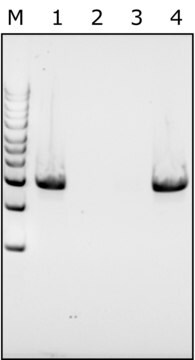
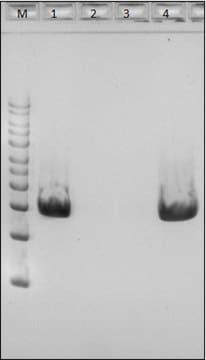
General description
Application
Biochem/physiol Actions
Components
- Achromopeptidase
- Chitinase
- Lyticase
- Lysostaphin
- Lysozyme
- Mutanolysin
Unit Definition
Signal Word
Danger
Hazard Statements
Precautionary Statements
Hazard Classifications
Resp. Sens. 1
Storage Class Code
11 - Combustible Solids
WGK
WGK 3
Flash Point(F)
Not applicable
Flash Point(C)
Not applicable
Regulatory Listings
Regulatory Listings are mainly provided for chemical products. Only limited information can be provided here for non-chemical products. No entry means none of the components are listed. It is the user’s obligation to ensure the safe and legal use of the product.
JAN Code
MAC4L-BULK:
MAC4LPROC:
MAC4L-5MG-PW:
MAC4L-5MG:
MAC4L-PH:
MAC4L-VAR:
Certificates of Analysis (COA)
Search for Certificates of Analysis (COA) by entering the products Lot/Batch Number. Lot and Batch Numbers can be found on a product’s label following the words ‘Lot’ or ‘Batch’.
Already Own This Product?
Find documentation for the products that you have recently purchased in the Document Library.
Customers Also Viewed
Articles
Enzymatic cell lysis and protoplast prep processes demystified, revealing complex enzymatic reactions during sample preparation.
Our team of scientists has experience in all areas of research including Life Science, Material Science, Chemical Synthesis, Chromatography, Analytical and many others.
Contact Technical Service