SRP0125
LSD1 substrate (Di-methylated K4_H3)
≥90% (HPLC), aqueous solution
Sinonimo/i:
HIST1H3E, Histone H3.1
Autenticatiper visualizzare i prezzi riservati alla tua organizzazione & contrattuali
About This Item
Codice UNSPSC:
12352204
NACRES:
NA.32
Prodotti consigliati
product name
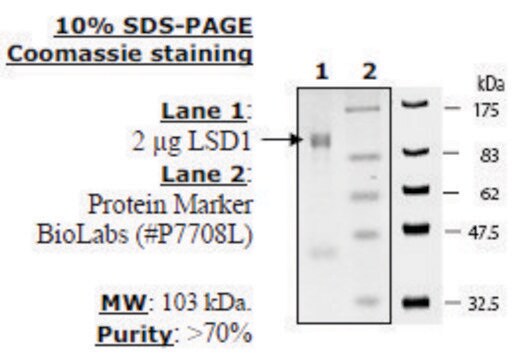
LSD1 substrate (Di-methylated K4_H3), ≥90% (HPLC)
Origine biologica
human
Saggio
≥90% (HPLC)
Forma fisica
aqueous solution
PM
2283 Da
Confezionamento
pkg of 500UL
Concentrazione
0.55 mg/mL
N° accesso UniProt
Condizioni di spedizione
dry ice
Temperatura di conservazione
−70°C
Informazioni sul gene
human ... HIST1H3E(8353)
Descrizione generale
Lysine-specific demethylase 1 (LSD1) acts on mono- and dimethylated histones and demethylates them. It has a role in embryogenesis, cell proliferation, spermatogenesis, adipogenesis and chromosomal segregation. Histone H3 peptide methylated at Lysine 4 is a substrate for LSD1. Histone cluster 1 H3 family member E (HIST1H3E) is a variant of histone 3 and is also called as the replicative histone. It is mainly expressed in the S-phase of the cell cycle. The gene encoding this protein is localized on human chromosome 6p22.
Applicazioni
Study enzyme kinetics and screen small molecular inhibitors of LSD1 for drug discovery and HTS applications
Azioni biochim/fisiol
Histones have an important role in the organization and modification of chromatin. Histone cluster 1 H3 family member E (HIST1H3E) has a role in DNA synthesis during DNA replication and might also be involved in DNA repair. It is deposited at the chromatin at the time of DNA replication-associated chromatin assembly. During herpes simplex virus 1 (HSV-1) infection, differential movement of HIST1H3E is associated with the assembly of viral chromatin.
Stato fisico
Supplied as 500 uL of water solution at 200 uM concetration.
Nota sulla preparazione
Thaw on ice. Upon first thaw, briefly spin tube to recover full content of the tube. Aliquot into single use aliquots. Store remaining product in aliquots at -70°C. Note: Avoid freeze/thaw cycles.
Codice della classe di stoccaggio
10 - Combustible liquids
Punto d’infiammabilità (°F)
Not applicable
Punto d’infiammabilità (°C)
Not applicable
Certificati d'analisi (COA)
Cerca il Certificati d'analisi (COA) digitando il numero di lotto/batch corrispondente. I numeri di lotto o di batch sono stampati sull'etichetta dei prodotti dopo la parola ‘Lotto’ o ‘Batch’.
Possiedi già questo prodotto?
I documenti relativi ai prodotti acquistati recentemente sono disponibili nell’Archivio dei documenti.
Yingwei Chen et al.
Critical reviews in eukaryotic gene expression, 22(1), 53-59 (2012-02-22)
Lysine-specific demethylase 1 (LSD1), the first identified histone demethylase, was belonged to the superfamily of the flavin adenine dinucleotide (FAD)-dependent amine oxidases. LSD1 specifically demethylates mono- or dimethylated dimethylated histone H3 lysine4 (H3K4) and H3 lysine 9 (H3K9) via a
Kami Ahmad et al.
Molecular cell, 9(6), 1191-1200 (2002-06-28)
Two very similar H3 histones-differing at only four amino acid positions-are produced in Drosophila cells. Here we describe a mechanism of chromatin regulation whereby the variant H3.3 is deposited at particular loci, including active rDNA arrays. While the major H3
Kristen L Conn et al.
PLoS pathogens, 9(10), e1003695-e1003695 (2013-10-17)
During lytic infections, HSV-1 genomes are assembled into unstable nucleosomes. The histones required for HSV-1 chromatin assembly, however, are in the cellular chromatin. We have shown that linker (H1) and core (H2B and H4) histones are mobilized during HSV-1 infection
Eric I Campos et al.
Nature structural & molecular biology, 17(11), 1343-1351 (2010-10-19)
The mechanism by which newly synthesized histones are imported into the nucleus and deposited onto replicating chromatin alongside segregating nucleosomal counterparts is poorly understood, yet this program is expected to bear on the putative epigenetic nature of histone post-translational modifications.
Alejandra Loyola et al.
Molecular cell, 24(2), 309-316 (2006-10-21)
Histone posttranslational modifications (PTMs) and sequence variants regulate genome function. Although accumulating evidence links particular PTM patterns with specific genomic loci, our knowledge concerning where and when these PTMs are imposed remains limited. Here, we find that lysine methylation is
Il team dei nostri ricercatori vanta grande esperienza in tutte le aree della ricerca quali Life Science, scienza dei materiali, sintesi chimica, cromatografia, discipline analitiche, ecc..
Contatta l'Assistenza Tecnica.